|
4/6/2023 0 Comments Protein backbone 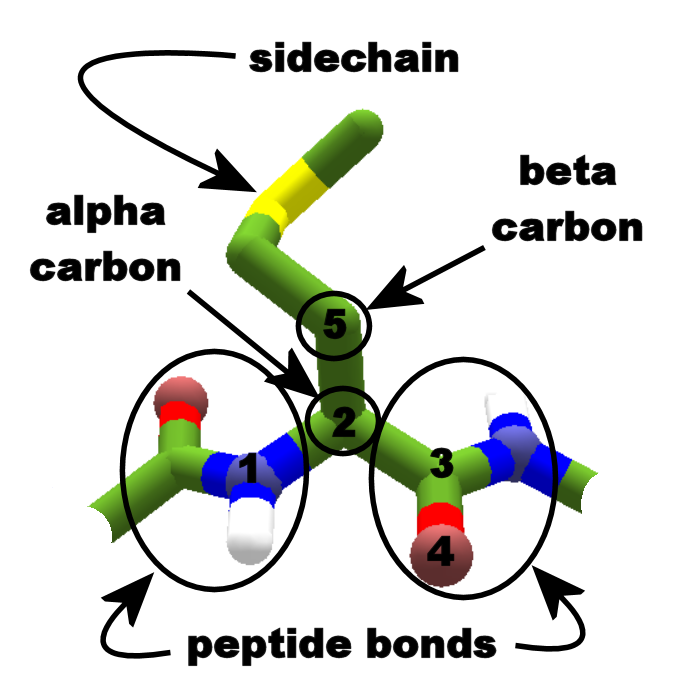
Thanks to a dedicated structure database, namely PTM-SD, a large screen of PTMs have been done and analyzed at local protein conformation levels using the structural alphabet protein blocks (PBs). Therefore, we analyzed the impact of the residue modifications on the protein structures at the local level. Nonetheless, the high diversity of PTM types and the multiple schemes of protein modifications (multiple PTMs, of different types, at different time, etc.) make difficult the direct confrontation of PTM annotations and protein structure data. Their impact on protein structures and their link to disorder regions have already been spotted in the past decade. modifications (PTMs) are known to play a critical role in the regulation of protein functions. 10 Laboratoire d'Excellence GR-Ex, 75739, Paris, France. 9 Institut National de la Transfusion Sanguine (INTS), 75739, Paris, France. 8 Université de Paris, Université de la Réunion, l'Université des Antilles, UMR_S 1134, 6, rue Alexandre Cabanel, 75739, Paris Cedex 15, France. 7 INSERM UMR_S 1134, BIGR, DSIMB, 75739, Paris, France.6 Department of Computer Science and Mathematics, Lebanese American University, Byblos, 1h401 2010, Lebanon.5 Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, USA.4 Laboratoire d'Excellence GR-Ex, 75739, Paris, France.3 Institut National de la Transfusion Sanguine (INTS), 75739, Paris, France.2 Université de Paris, Université de la Réunion, l'Université des Antilles, UMR_S 1134, 6, rue Alexandre Cabanel, 75739, Paris Cedex 15, France.1 INSERM UMR_S 1134, BIGR, DSIMB, 75739, Paris, France.Secondary structure which has links to other pages with details on alpha helices, beta sheets, and turns.Introduction to molecular visualization.Notice how the arrowheads point towards the C terminus. (Proteins are synthesized by adding amino acids to the C terminus.) This color scheme is called N->C Rainbow. A such a ribbon representation is with a spectral sequence of colors from the amino (N) terminus to the carboxy (C) terminus.Arrowheads point towards the carboxy terminus. This type of representation is properly called a secondary structure schematic, but is called a cartoon in Jmol and its family of ancestral visualization programs ( RasMol, Chime). The helices and strands are represented as ribbons, while the "ropes" connecting them are smoothed backbone traces. This domain contains the alpha helix used above, but also contains a small beta sheet made of four beta strands, plus loops (regions that are neither alpha helix nor beta strand) connecting the helices and strands. The arrowhead at one end points to the carboxyl terminus. As you can see, the ribbon is a smoothed backbone trace expanded in width. Here the ribbon is violet , the standard secondary structure color for alpha helices. Perhaps the most common protein backbone representation is the. Here, the smoothed backbone trace is green . Note that the backbone trace does not follow any actual covalent chemical bonds - it simplyĬonnects alpha carbon positions, thereby simplifying the representation.Ī is another common backbone representation. : Now we'll draw a yellow line between alpha carbons (balls). We could also, leaving only the atoms that are part of the main chain, also called the backbone. Each amino acid's main chain atoms are N-C-C, where the first C is the alpha carbon (shown as a ball), and the second, the carboxyl carbon with its double-bonded oxygen (double bonds not shown). Hydrogen atoms make up almost exactly 50% of the atoms in proteins. Lets begin with (15 amino acids) The atoms and bonds are colored by element: 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
